-
Call Now
1800-102-2727
Restriction Enzymes, Discovery, Mechanism of Action, Nomenclature, Types, Importance, Drawbacks, Practice problems and FAQs
It is very fun to do some DIY (do it yourself) crafts if you are really interested in it. You might have done it in your art and craft classes. Cutting colour papers and pasting them on each other and making creative crafts are hobbies of many people. It is beautiful to see those crafts after work, right? So can you tell me an important tool required for such hand craft work? Yes, scissors. Without scissors we cannot make the craft.
Do you know there are scissors used in the process of biotechnology also? We can call them molecular scissors. These will do the same work as scissors. These are used to cut the DNA molecules near or at certain sequences called restriction sites.
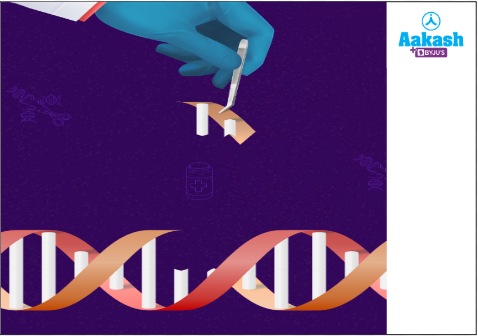
Fig: Action of restriction endonucleases
Restriction endonucleases are commonly used in the transformation, transfection, and other related processes as a tool for gene modification. But how do they work and what is their role in the gene cloning process?
It is an interesting topic to discuss, right? So let’s discuss restriction enzymes in depth in this article and find out the answers for all the above questions.
Table of contents:
- Restriction enzymes
- Nucleases
- Discovery of restriction endonucleases
- Recognition sequences
- Mechanism of action of restriction endonucleases
- Nomenclature of restriction endonucleases
- Insertion of GOI into the plasmid
- Types of restriction enzymes
- Common restriction enzymes
- Importance of restriction enzymes
- Drawbacks of restriction enzymes
- Practice problems
- FAQs
Restriction endonucleases
Restriction endonucleases are enzymes which cleave the DNA by digesting phosphodiester bonds at a specific sequence called recognition sequence within the DNA. Examples include EcoR I. The word ‘restriction’ in restriction enzyme refers to prevention of the multiplication of bacteriophages in bacteria.
If we isolate a DNA with GOI (gene of interest), then the next step is to transfer it into the host organism. Here a vector DNA is helping to transfer the GOI to the host. But how will the DNA with GOI get inserted into the cloning vector? It cannot be simply inserted. For that we need to cut open the DNA plasmid. In this step of recombinant DNA technology we will use restriction endonuclease or REN.
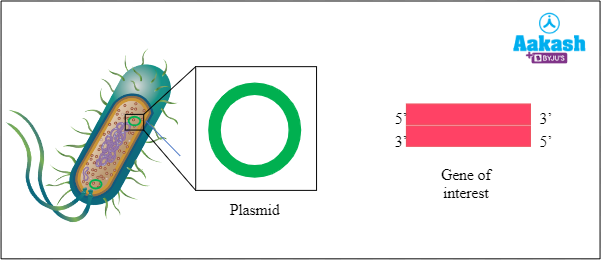
Fig: Plasmid and GOI
Restriction Enzymes are the workhorses of recombinant DNA technology. They are one of the most important tools in RDT.
Most common enzymes
You may know about the proteases which break down the proteins, lipases which break down the lipids and nucleases which cleave nucleic acids. Right? All these proteases, lipases and nucleases are the enzymes which have the ability to break molecules. Restriction endonucleases belong to the family of nucleases. REN can break the phosphodiester bonds in the DNA and RNA.
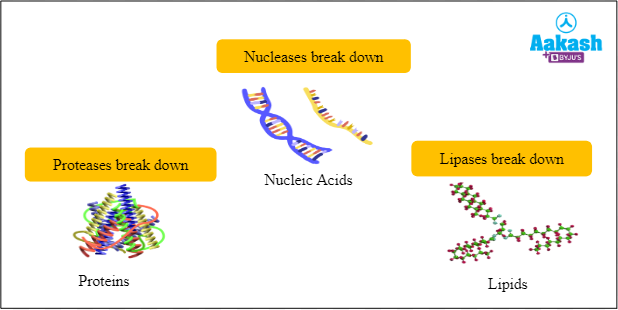
Fig: Types of enzymes
Nucleases
Any enzyme that helps in the cleavage of nucleic acids is called nucleases. They are able to cut the DNA into fragments by breaking the phosphodiester bonds.
Classification of nucleases
Depending on the actions of nucleases, they can be divided into two. They are the endonucleases and the exonucleases.
Fig: Classification of nucleases
Endonucleases
Endo means inside, hence endonucleases are enzymes that cleave DNA at any point within the DNA strand but not at the ends. They act within the polynucleotide chain.
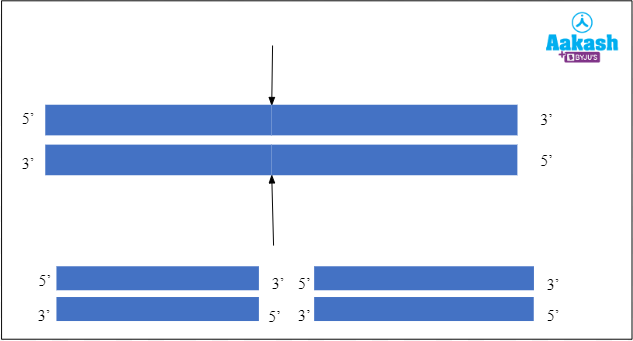
Fig: Mechanism of action of endonuclease
Exonucleases
Exo means outside, hence exonucleases are enzymes that cut either the double stranded or single stranded DNA and remove the nucleotides from the 5’ or 3’ ends. They act on the ends of the polynucleotide chain. They can act either on the 3’ or the 5’ ends and remove the nucleotides one by one at a time.
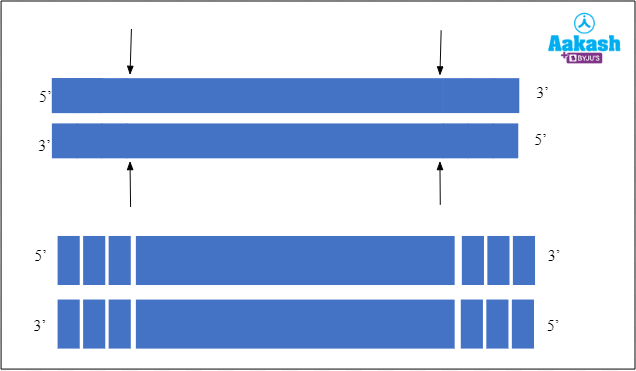
Fig: Mechanism of action of exonuclease
Discovery of restriction endonucleases
Restriction endonucleases are enzymes found in certain bacteria that cut the DNA molecule at or near specific nucleotide sequences. Bacteriophages or phages are the viruses that attack and infect bacterial cells. These viruses attach to the surface of bacterial cells and inject their genetic material into the bacterial cell. They then use the machinery of bacterial cells to multiply and destroy them at the time of release of new phage particles.
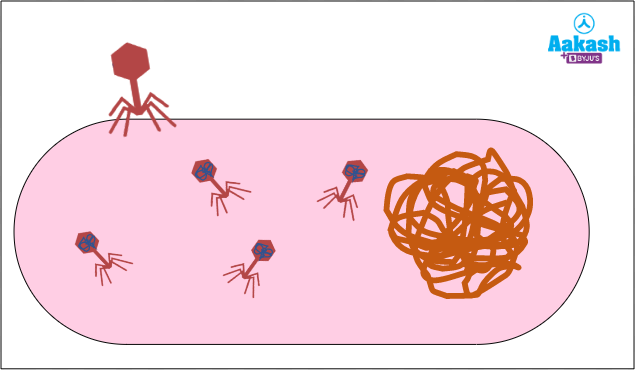
Fig: Bacteriophage attacking bacteria and using the bacterial cell machinery for multiplication
Restriction modification system in bacteria
Many bacteria are susceptible to these viruses. Restriction endonucleases were discovered in such bacteria and are a part of the defence mechanism called the restriction modification system against attack by bacteriophages. Two types of enzymes are found in bacteria in this defence mechanism. One is the restriction enzymes and the other is the modification enzymes.
Fig: Enzymes of restriction modification system in bacteria
Restriction enzymes
These can recognise the foreign DNA molecule and cut up the foreign bacteriophage DNA. In this way these enzymes restrict the propagation of foreign DNA in the bacterium. So that bacteriophage cannot multiply inside the bacteria.

Fig: Mechanism of action of restriction enzyme on the foreign phage DNA
Modification enzymes
The second enzyme is the modification enzyme that recognises and modifies the bacterial DNA to protect it from the DNA-degrading activity of its very own restriction enzyme. The modification enzymes modify the bacterial genome by adding methyl groups to one or more bases within the recognition sequence of restriction enzymes. The DNA with the modified sequences cannot be recognised and cut by restriction enzymes. In this way, this modification prevents the bacterial genome getting attacked by the restriction enzyme. The restriction modification system was first discovered in the bacterium Escherichia coli in late 1960s.
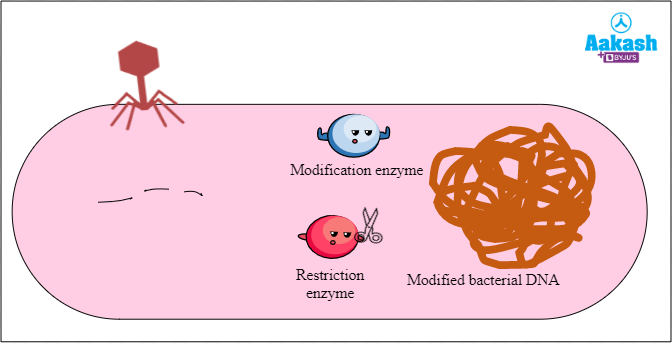
Fig: Action of modification enzyme
Recognition sequences or recognition sites
Recognition sequence is the site where the DNA is cut by a restriction endonuclease. These are palindromic sequences. For example, EcoR I recognises the palindromic sequence 5' - GAATTC - 3'. More than 900 restriction enzymes that have been isolated from over 230 strains of bacteria and each recognises different recognition sequences.
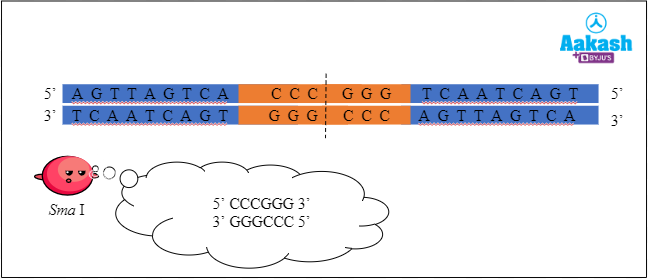
Fig: Restriction enzyme Sma I cutting the DNA at the recognition sequence
Palindromic sequences
Have you heard about palindromes in English? These are the words which read the same when we read the letters from left to right and right to left. For example let’s take the word ‘MALAYALAM’. Now if we start reading this word backwards, it will still read the same.
Similar to the palindromes in the English language, the recognition sequences of restriction enzymes read the same forward and backward, but in different strands. Hence such a sequence is called a palindromic sequence.
We usually read the nucleotides on a DNA in the 5’🡪 3’ direction. When the bases read the same on both the strands from 5’🡪 3’ direction, the sequence is said to be palindromic. Palindromic sequences are nucleotide sequences that read the same on the two strands when orientation of the reading is kept the same, that is, 5'→3' direction. For example, GAATTC from 5’ to 3’ on one DNA strand and CTTAAG from 3’ to 5’ in the complementary DNA strand is a palindromic sequence.
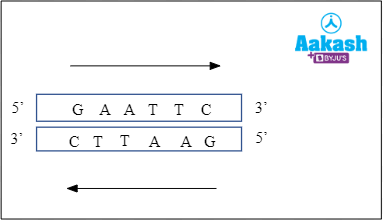
Fig: Palindromic sequence of EcoR I restriction endonuclease
Now it is clear that the restriction enzymes will be making cuts on the palindromic sequences which will be the recognition sites. But how is it doing? So next we are going to discuss the mechanism of action of restriction enzymes.
Mechanism of action of restriction endonucleases
Restriction enzymes cut the DNA into fragments of a size suitable for cloning. The REN produce two types of cuts in DNA as follows:
- Blunt cuts that result in non-cohesive or blunt ends.
- Staggered cuts that result in cohesive or sticky ends.

Fig: Types of cut by restriction endonucleases
Blunt ends
These are formed when the restriction endonucleases cut the DNA strands in such a way that both strands terminate in a base pair. There are no unpaired nucleotides at the end of the DNA molecule. Here the restriction enzymes cut exactly at the middle of the recognition sequence to form blunt ends.
For example, Sma I is the restriction enzyme obtained from Serratia marcescens. It recognises the palindromic sequence 5' - CCCGGG - 3' and makes the following cut between C and G on both the strands of DNA to form blunt ends.
5' - CCC↓GGG - 3'
3' - CGG↑CCC - 5'
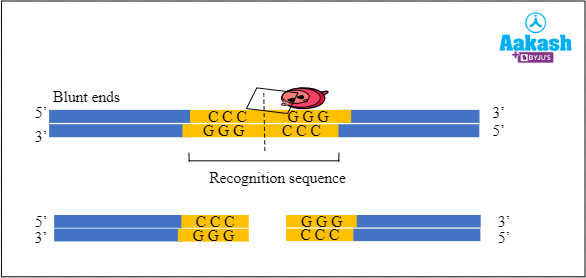
Fig: Formation of blunt ends by the restriction endonuclease Sma I
Sticky ends
It is a fragment of DNA produced by a staggered cut using restriction enzymes, in which the terminal portion has a stretch of unpaired nucleotides. Formation of sticky ends is done as follows:
- Restriction endonucleases break the phosphodiester bond between two specific adjacent nucleotides of a strand.
- The restriction enzyme cuts DNA at one end of the sequence, between two bases on the same strand, then cuts on the opposite end of the complementary strand in the same way.
- This results in the formation of two single stranded ends of DNA containing few nucleotides without any complementary bases. These ends are called sticky ends.
- When complementary sticky ends of DNA fragments meet, hydrogen bonds are formed between the complementary bases which results in annealing of the ends of the DNA fragments.
For example, EcoR I is the restriction enzyme obtained from Escherichia coli. It recognises the palindromic sequence 5' - GAATTC - 3' and makes the following cut between G and A on both the strands of DNA to form sticky ends.
5' - G↓AATTC - 3'
3' - CTTAA↑G - 5'
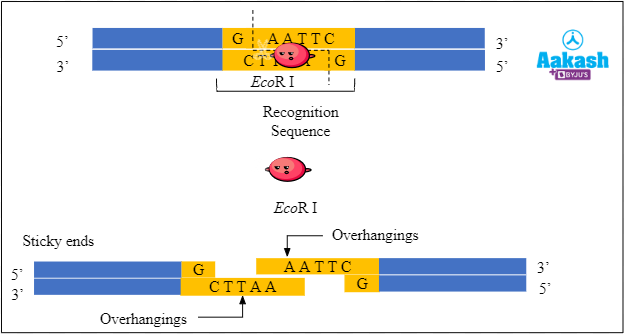
Fig: Formation of sticky ends by the restriction endonuclease EcoR I
Common restriction endonucleases and the types of cuts
Different restriction endonucleases make different types of cuts. Now let’s see the different types of cut made by the common restriction endonucleases.
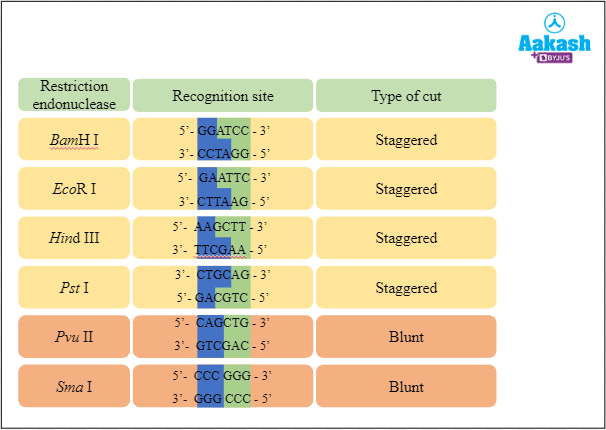
Fig: Types of cuts made by the common restriction endonucleases
Nomenclature of restriction endonucleases
Restriction endonucleases are named according to the organism from which they were discovered, using a system of letters and numbers. The nomenclature follows certain rules while naming the restriction endonuclease. The rules are as follows:
- The 1st part should be the first letter of the genus name of the bacteria from which it is discovered.
- The 2nd part should be the first 2 letters of the species name of the bacteria.
- The 3rd part should be the strain of the bacteria.
- The 4th part should be the order of discovery in Roman numerals.
For example, we will see the naming of the restriction endonuclease EcoR I. EcoR I was isolated from Escherichia coli RY 13 strain. The first letter comes from the genus name ‘Escherichia’, the next 2 letters coming from the species name ‘coli’. The third letter comes from the strain of the bacteria ‘RY 13’. Lastly, the Roman numerals are used to identify specific enzymes from bacteria that contain multiple restriction enzymes indicating the order in which restriction enzymes were discovered from a particular strain.
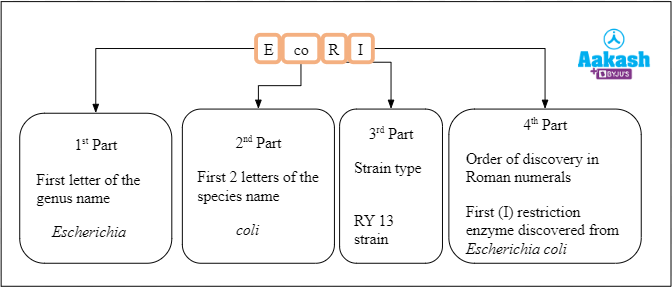
Fig: Nomenclature of the restriction endonuclease EcoR I
Insertion of GOI (gene of interest) into plasmid
Till now we have seen what restriction enzymes are, where do they present, how do they function, how are they named and different types of REN. Next, what we have to do is to insert the GOI into the plasmid to produce rDNA. Let’s see how it is done.
Creation of suitable gene of interest (GOI)
Before the action of REN, the DNA with GOI has to undergo PCR. After the first PCR we have to do another PCR to add some extra flanking sequences on the side. This GOI DNA is now ready to be inserted into the plasmid.
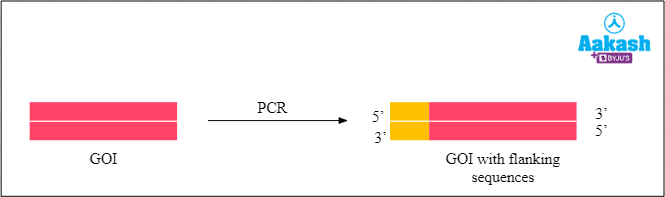
Fig: Final GOI
Cutting of pBR322 plasmid
Here in this case we are going to take pBR322 as the plasmid. This plasmid has an EcoR I recognition site. So we are going to use the EcoR I restriction enzyme on it.
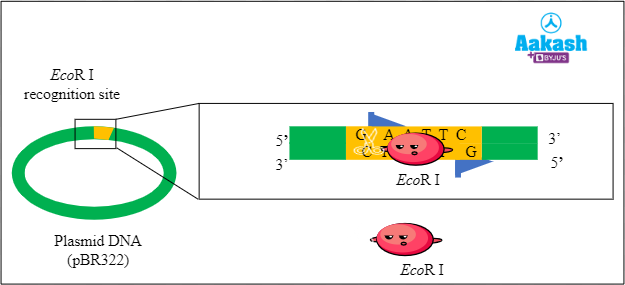
Fig: Digestion of plasmid with the restriction endonuclease
First, the plasmid is digested or cut with the restriction enzyme EcoR I. This step produces fragments with sticky ends.
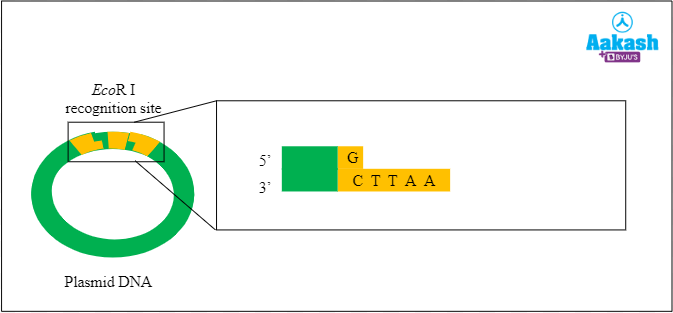
Fig: Fragmented plasmid
Cutting of gene of interest (GOI)
We will do the same for the GOI as well. The gene fragment is digested or cut with the same restriction enzyme EcoR I. This is done to join the GOI with the plasmid.
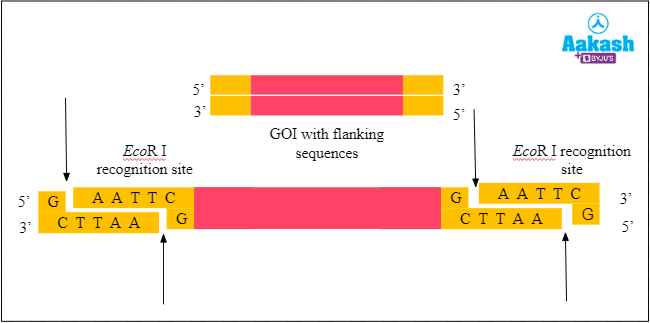
Fig: Digestion of GOI
This step produces fragments with sticky ends. All of the ends have an overhang of four nucleotides, with the sequence 5’- AATT - 3’. So let’s remove those fragments.
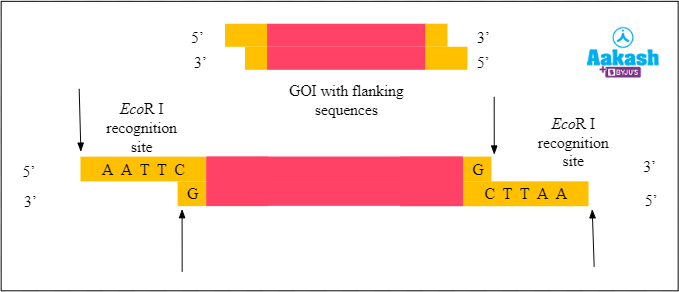
Fig: GOI with sticky ends
Creation of chimeric DNA
Now we have both the plasmid and the GOI with sticky ends. Next we have to join them together. The sticky ends of the two fragments (plasmid and GOI) stick together by complementary base pairing. It results in the formation of chimeric DNA.
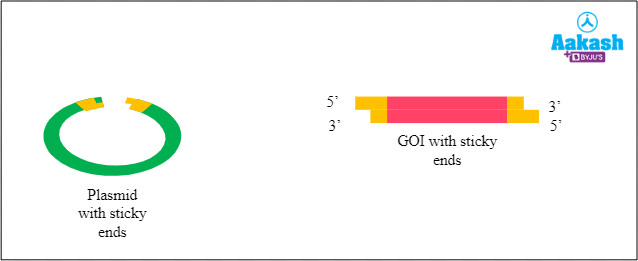
Fig: Formation of chimeric DNA
Mechanism of action of ligase enzyme
Ligase enzymes catalyse the formation of phosphodiester bonds, thus joining the desired DNA fragments. In genetic engineering it helps in joining the vector DNA with the DNA of interest to form the rDNA. There are still gaps in the sugar-phosphate backbones of the DNA double helix at the joining sites where the gene of interest (GOI) and plasmid DNA meet. So the ligase enzyme is used to complete the process by ligating the 2 DNA strands in the below given manner:
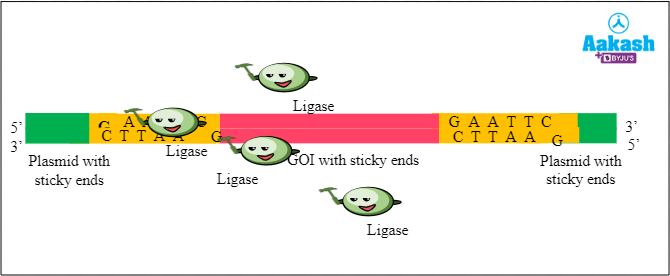
Fig: Mechanism of action of ligase enzyme
Recombinant DNA
Now we have the GOI inserted into the plasmid. This is the way we get the recombinant DNA, rDNA or chimeric DNA.
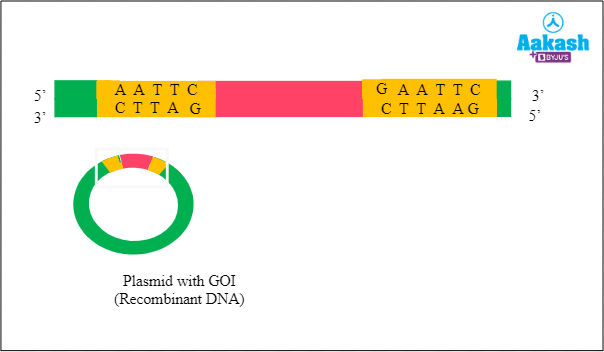
Fig: Recombinant DNA
Types of restriction enzymes
The restriction enzymes can be classified into four types; type I, II, III and IV.
Factors deciding classification of restriction endonuclease
This classification is done on the basis of the following factors:
- Subunit composition - It is a complex or simple enzyme. For example, some restriction endonucleases possess restriction and modification enzymes.
- Cleavage position - The enzyme is performing the cleavage near, within or far from the recognition sequence.
- Sequence specificity - The type of sequence involved.
- Cofactor requirements - Some restriction endonucleases require cofactors for their action. For example, Hind III requires the presence of cofactor Mg2+ for its action.
Type I enzymes
These are complex, multisubunit enzymes with a combination of restriction and modification enzymes. It can cut DNA at random sites far from their recognition sequences. They can not produce discrete restriction fragments, hence have very little practical value in recombinant DNA technology.
Type II enzymes
They can cut the DNA at defined positions. The cleavage made by the type II enzymes will be within their recognition sequences and produce discrete restriction fragments. They are the group of enzymes predominantly used in gene cloning. They are a collection of unrelated proteins. Most common type II enzymes are Hind III and Not I.
Type III Enzymes
The cleavage of type III enzymes is outside their recognition sequences. They also possess restriction and modification enzymes. Hence to accomplish the cleavage, two recognition sequences are required in the opposite orientation in the same DNA. The complete digestion of sequences is rare with this enzyme.
Type IV Enzymes
Modified or methylated DNA is recognised specifically by Type IV enzymes. Their sequence specificity is also very weak.
Common restriction enzymes
Now we will discuss some of the common restriction enzymes used for recognising the specific DNA sequences and cleaving them. BamH I, Pvu I, Cla I, Hind III, Hind II, Pst I, Pvu II, EcoR I, Xho I, Not I, Hae III and Sal I are some of the common restriction endonucleases.
BamH I
This restriction enzyme is isolated from the bacterium Bacillus amyloliquefaciens. It is a type II restriction enzyme. The recognition sequence where the BamH I binds is 5'- GGATCC -3'. It can cleave the recognition sequences just after the 5'- guanine on each strand. Like other type II enzymes, BamH I also needs divalent metals as cofactors and it helps in cleaving the DNA in the following way:
5' - G↓GATCC - 3'
3’- CCTAG↑G - 5’
Pvu I
This restriction enzyme is isolated from the bacterium Proteus vulgaris. The restriction sequence of this enzyme is 5′ - CGATCG - 3′. It will generate fragments with 3′ - cohesive termini. It can cleave the recognition sequences just after the thymine on each strand in the following way:
5′ - CGAT↓CG - 3′
3’ - GC↑TAGC - 5’
Cla I
The restriction enzyme isolated from Caryophanon latum L. is called Cla I. It cleaves the recognition sequence 5′ - ATCGAT - 3′ of the DNA and it generates DNA fragments with 5′- cohesive ends. It can cleave the recognition sequences just after the 5' - thymine on each strand. If the DNA isolated from Escherichia coli has the dam methylase (dam+ strains), then Cla I will only cleave the DNA partially.
5′- AT↓CGAT - 3′
3’- TAGC↑TA - 5’
Hind III
The restriction enzyme isolated from Haemophilus influenzae is the Hind III. It is a type II restriction enzyme. The cleavage of the palindromic sequence 5’- AAGCTT - 3’ is done by Hind III. It can cleave the recognition sequences just after the 5' - adenine on each strand. It happens in the presence of cofactor Mg2+. It also produces sticky ends after the cleavage in the following way:
5' - A↓AGCTT - 3'
3’ - TTCGA↑A - 5’
Hind II
It is the first type II restriction enzyme identified. It is isolated from Haemophilus influenzae. Hind I was identified after Hind II, but isolated first. Hind II got the number ‘II’ , because it was isolated after the Hind I enzyme. Hind II generates blunt ends by recognising a specific sequence of 6 base pairs, 5’- GTCGAC - 3’and makes a cut in the DNA in the following way:
5’- GTC↓GAC -3’
3’- CAG↑CTG -5’
Pst I
It is isolated from Providencia stuartii, which is a gram-negative bacteria. It is a type II enzyme.
The cleavage site is the recognition sequence 5′ - CTGCAG - 3′. It can cleave the recognition sequences just after the 3' - adenine on each strand. The fragments generated have 3′ - cohesive termini and yield sticky ends in the following way:
5′- CTGCA↓G -3′
3’- G↑ACGTC -5’
Pvu II
This restriction enzyme is isolated from the bacterium Proteus vulgaris, which is a gram-negative bacteria. The restriction sequence of this enzyme is 5' - CAG/CTG - 3'. It can cleave the recognition sequences just after the 5' - guanine and cytosine on each strand in the following way:
5' - CAG↓CTG - 3'
3’ - GTC↑GAC - 5’
EcoR I
The restriction enzyme isolated from the Escherichia coli bacteria RY 13 strain is called EcoR I. The recognition sequence where it makes cleavage is 5’ - G/AATTC - 3’. It can cleave the recognition sequences just after the 5' - guanine and adenine on each strand. It makes sticky ends with 5’ overhangings in the following way:
5’ - G↓AATTC - 3’
3’ - CTTAA↑G - 5’
Sal I
The restriction enzyme isolated from Streptomyces albus is called Sal I. It cleaves the recognition sequence 5’ - G/TCGAC - 3’ of the DNA. It can cleave the recognition sequences just after the 5' - guanine and thymine on each strand. It generates fragments with 5’ cohesive ends in the following way:
5’ - G↓TCGAC - 3’
3’ - CAGCT↑G - 5’
Not I
The restriction enzyme isolated from the bacterium Nocardia otitidis is called Not I. It creates sticky ends by cutting the restriction sequence 5’- GC/GGCCGC - 3’. It is included in the class of ‘rare-cutter enzymes’.
5’ - GC↓GGCCGC - 3’
3’ - CGCCGG↑CG - 5’
Hae III
The third endonuclease isolated from the bacteria Haemophilus aegyptius is called Hae III. it cleaves the DNA at the site of 5′ - GG/CC - 3’. Blunt ends are formed in the fragments created after the cut. This enzyme is not very effective for making cleavage in single stranded DNA.
5′ - GG↓CC - 3’
3’ - CC↑GG - 5’
Xho I
Restriction enzymes Xho I, is obtained from Xanthomonas holcicola. It produces sticky ends by making the following cut in the recognition sequence:
5 ' - C↓TCGAG - 3'
3’ - GAGCT↑C - 5’
Importance of restriction endonucleases in recombinant DNA technology
Importance of REN in RDT is as follows:
- The discovery of restriction endonucleases in the 1960s helped in the making of the first artificial recombinant DNA in 1972 by Cohen and Boyer.
- These enzymes catalyse the cleavage of DNA and thus help in isolating the gene of interest.
- It also helps in creating the sticky ends in vectors and DNA of interest for the formation of rDNA. The most important step in genetic engineering is the formation of recombinant DNA (rDNA). The formation of the recombinant DNA depends on the use of restriction enzymes.
- The discovery and the availability of pure forms of restriction endonucleases is therefore an important milestone in the development of genetic engineering.
Drawbacks of restriction endonucleases
The following are the major drawbacks of restriction endonucleases:
- Restriction endonucleases may lose their activity due to improper handling or storage. This will result in the incomplete or no digestion of DNA.
- Use of too much or too little restriction enzymes will result in the incomplete digestion of DNA.
- Sometimes restriction enzymes may show star activity or off-target cleavage where these will cleave sites that are similar, but not identical to the recognition sequences.
- If restriction enzymes, cut at random sites then vector DNA and DNA of interest would get fragmented making it unfit for forming rDNA.
Practice Problems
1. How many of the statements below are true?
I. If a linear DNA molecule is digested with a restriction enzyme at four recognition sites, 4 fragments would be produced.
II. Six fragments are formed when a closed circular DNA molecule is digested by a restriction enzyme at six recognition sites.
III. If a linear DNA molecule is digested with a restriction enzyme at four recognition sites on the DNA, 5 fragments would be produced.
IV. If a closed circular DNA molecule is digested with a restriction enzyme at six recognition sites, 7 fragments would be produced.
- 1
- 2
- 3
- 4
Solution: Restriction endonucleases are the enzymes that cleave the phosphodiester bonds present between the consecutive nucleotides within or near to specific DNA sequences that they identify. These identifiable sequences by restriction endonucleases are known as recognition sequences. If a linear structure is cut into fragments, the number of fragments is always one greater than the number of cuts being done in it. If ‘n’ is the number of cuts made in the linear structure, then ‘n+1’ would be the number of fragments produced. When a closed circular DNA is cut into fragments, the number of fragments produced is always the same as the number of cuts made. If ‘n’ is the number of cuts made in the closed circular structure, ‘n’ would be the number of fragments produced. Here, the number of recognition sites signifies the number of cuts being done by the restriction enzymes. If a linear DNA molecule is digested with a restriction enzyme at four recognition sites, 5 fragments would be produced. Six fragments are formed when a closed circular DNA molecule is digested with a restriction enzyme at six recognition sites. Hence the statements II and III are true, but statements I and IV are false. So the correct option is b.
2. In an experiment, the genomic DNA was isolated from three different eukaryotic sources, A, B and C. The genes of interest from all three sources were amplified. These were subjected to digestion with one restriction enzyme and then electrophoresed in agarose gel. Provided that the restriction endonuclease used cleaved the DNA within the recognition sequence, study the figure given below carefully and match the options in column I with that of column II.
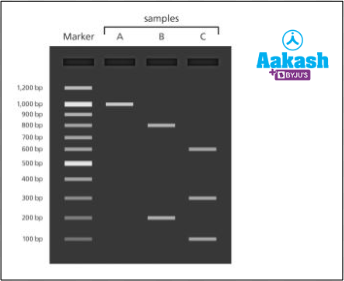
Fig: DNA fragments on agarose gel
|
Column I (DNA sample) |
Column II (Number of recognition sites) |
||
|
I |
DNA of sample A |
p |
Contains two recognition sites |
|
II |
DNA of sample B |
q |
Does not contain any recognition site |
|
III |
DNA of sample C |
r |
Contains one recognition site |
- I - r, II - q, III - p
- I - q, II - r, III - p
- I - q, II - p, III - r
- I - q, II - p, III - r
Solution: The gene of interest obtained from source A has been found to form one band in the lane corresponding to the well where it was loaded. Hence, it is intact even after treatment with the restriction enzyme. So, DNA of sample A did not contain any recognition site. The gene of interest isolated from source B has been found to form two bands in the lane corresponding to the well where it was loaded. Hence, it can be assumed to have one recognition site. Treatment with the restriction enzyme must have cleaved it into two pieces. The gene of interest from source C, after restriction digestion, has been found to form three bands in the lane corresponding to the well where it was loaded. Hence, it can be assumed to have two recognition sites. So treatment with the restriction enzyme must have cleaved it into three pieces. Hence the correct option is b.
3. Assertion: Restriction enzymes recognise specific palindromic sequences.
Reason: Palindromic sequences are DNA sequences that are read the same way when the orientation is kept the same in both the strands.
- Both the assertion and the reason are true and the reason is the correct explanation of the assertion
- Both the assertion and the reason are true but the reason is not the correct explanation of the assertion
- Assertion is true but the reason is false
- Both assertion and reason are false
Solution: Restriction enzymes are a type of endonucleases that cleave phosphodiester bonds between consecutive nucleotides within or near to specific DNA sequences known as recognition sites. Recognition sites are the sites on the polynucleotide strand that are usually palindromic in nature, identified by a restriction enzyme for the purpose of cleavage. Palindromic sequences are DNA sequences that are read the same way forward or backward when the polarity is kept the same in both the strands. For example, GAATTC from 5’ to 3’ on one DNA strand and CTTAAG from 3’ to 5’ in the complementary DNA strand is a palindromic sequence as shown below.
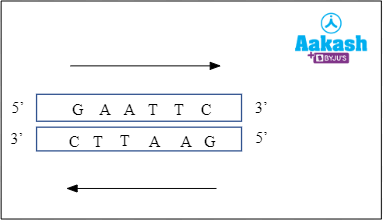
Fig: Palindromic sequence
Hence, both the assertion and the reason are true but the reason is not the correct explanation of the assertion. So option b is correct.
4. A person trying to create a recombinant DNA molecule failed to do so. Below is the procedure that he followed.
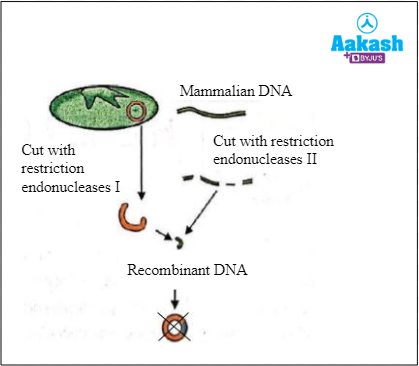
Fig: Steps of recombinant DNA technology followed
Which among the following is the most probable mistake he might have made?
- He did not use the enzyme DNA polymerase
- He used double stranded mammalian DNA
- He used two different restriction enzymes
- He inserted only one gene of the mammalian DNA into the plasmid
Solution: The mammalian DNA and the plasmid DNA should be cut by the same restriction endonuclease to obtain the complementary sticky ends in recombinant DNA technology. These enzymes cut the sugar phosphate backbone in the double stranded DNA at or near specific points called recognition sites that can be palindromic in nature. Here, in the question, the person has used two different restriction endonucleases. Therefore, he could not get the complementary sticky ends as a result of which he was not able to create rDNA. The correct procedure to create a rDNA where the mammalian DNA as well as plasmid DNA of bacteria are cut by the same restriction endonuclease is given below. The enzyme DNA polymerase catalyses the addition of nucleotides to the 3’ end of a polynucleotide, which is not required in rDNA technology. It is used in DNA replication. DNA used in rDNA technology needs to be double stranded. rDNA can be obtained by linking just one gene to the plasmid as well. Hence the correct option is c.
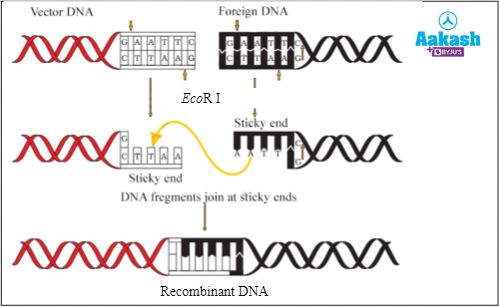
Fig: Formation of recombinant DNA using a single restriction endonuclease
5. How many sticky ends will be produced if a plasmid vector is digested at a single site by EcoR I?
- 1
- 2
- 4
- 6
Solution: Restriction endonucleases are the enzymes that cleave the phosphodiester bonds present between consecutive nucleotides within specific DNA sequences known as recognition sites. EcoR I is a restriction endonuclease. It was isolated from Escherichia coli RY13 bacterium. It recognises the 5' - GAATTC - 3' nucleotide sequence in a double stranded DNA and cuts between G and A. As the cut is made slightly away from the centre, it results in the formation of DNA fragments with single stranded overhanging free ends called sticky ends. If there are cuts between G and A in both the strands in the plasmid DNA, this results in the formation of a DNA fragment with two sticky ends. Hence, two sticky ends are produced if a plasmid vector is digested at a single site by EcoR I. Hence the correct option is b.
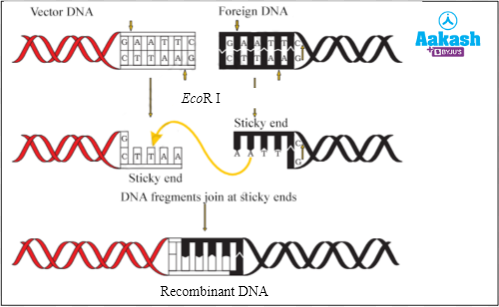
Fig: Mechanism of action of EcoR I
FAQs
1. Who was the first to discover restriction endonuclease?
Answer: W. Arber and H. Smith received the Nobel Prize in Medicine in 1978 for discovering restriction endonucleases, and D. Nathans for establishing that restriction endonucleases may be used to physically map DNA.
2. Which restriction endonuclease is isolated first?
Answer: The restriction endonuclease Hind II was the first to be discovered. It recognises the 6 base pairs long nucleotide sequences, 5’- GTCGAC - 3’and makes a cut in the DNA in the following way:
5’ - GTC↓GAC - 3’
3’ - CAG↑CTG - 5’
3. What are isoschizomers and neoschizomers?
Answer: Isoschizomers are the restriction enzymes which recognise and cleave at the same recognition site. For example, Sph I (CGTAC/G) and BbuI (CGTAC/G) are isoschizomers of each other. Neoschizomers are the restriction enzymes which recognise the same site and have a different cleavage pattern.
4. What is the difference between a linker and an adapter?
Answer: Linkers are oligonucleotides with two blunt ends that are chemically synthesised. It has two strands. Adaptors are oligonucleotides with one blunt end and one sticky end that have been chemically synthesised. Single-stranded and double-stranded versions are available.
5. What is homopolymer tailing, and how does it work?
Answer: The ligation of blunt-ended DNA fragments is accomplished through homopolymer tailing. The enzyme terminal deoxynucleotidyl transferase is used for tailing. This enzyme will add a nucleotide to the 3' end of a double-stranded DNA molecule on a regular basis.


