-
Call Now
1800-102-2727
Central Dogma, Transcription, Reverse Transcription Practice Problems and FAQs
We all have different types of hair. Right? It differs in their colour and texture. If you think
about the texture some will have straight hair, some have curly hair and others have wavy hair. If
you think about the colour some have brown hairs, some have black hairs and some have grey
hairs.

Fig: Difference in hair colour and texture
So what is the reason behind these variations in the structure of hair? Yes, it depends on the
hair follicles, enzymes and proteins like keratin, trichohyalin etc. You are familiar with the
keratin protein, which is a protein that makes the hair shine and strong. So the variation in the
amounts of keratin will also make a difference in the structure of hair. A single protein can
make this much change. Interesting, right? But why does the amount of protein differ in
different individuals? The reason is hidden inside their genes.
If we make it more scientific we can say that the proteins are synthesised using RNA by
ribosomes. Then where does RNA come from? It is synthesised from DNA through a process
called transcription. So how does RNA synthesis occur? You would like to know more about
this. So we are going to discuss the process of transcription in depth in this article.
Table of contents
- Central dogma
- Transcription
- Transcription in prokaryotes
- Transcription in eukaryotes
- Post transcriptional modifications
- Reverse transcription
- Significances of transcription and post transcriptional modifications
- Practice Problems
- FAQs
Central dogma
Central dogma of molecular biology was proposed by Francis Crick in September 1957. This
process involves the conversion of DNA into a functional product (protein). According to
central dogma the biological information flows from DNA to RNA and from there to
proteins.

Fig : Francis Crick
Process of central dogma
DNA is the blueprint of all the information. DNA recreates itself by the process of replication.
The RNA is formed from DNA by the process of transcription. Information in RNA is decoded
and translated into protein. The whole process of replication of DNA, transcription of DNA and
translation of RNA together is called central dogma.

Fig: Central dogma
Here in this article we are going to discuss the process of transcription where the RNAs are
formed from DNAs.
Transcription
The process of copying genetic information from DNA (deoxy ribo nucleic acid) into RNA
(ribonucleic acid) is called transcription.
We know that during replication, whole DNA is replicated to make a new copy. But in
transcription the information is transferred from one daughter strand to RNA (3’ → 5’). How
does this happen then? Let’s explore.
Direction of transcription
Enzymes are essential to carry out all the molecular processes. DNA dependent DNA polymerase is involved in the replication, because the enzyme uses DNA as a template. Similarly, transcription is carried out by an enzyme known as DNA dependent RNA polymerase. Similar to DNA polymerase, RNA polymerase also catalyses transcription in the direction 5’ to 3’.

Fig: Direction of transcription
Action of RNA polymerase
Unlike DNA polymerase, RNA polymerase does not require a primer to initiate RNA synthesis.
DNA dependent RNA polymerase adds Uracil instead of Thymine.
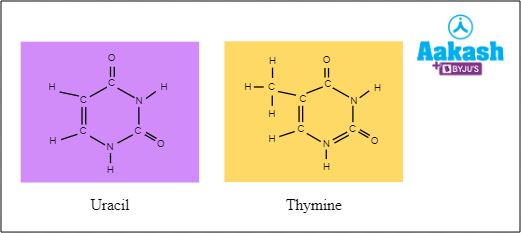
Fig: Structure of uracil and thymine
Coding and non coding strands
That segment of DNA getting transcribed is known as the transcription unit. During replication, both these strands get copied. Whereas in transcription, only one strand of DNA gets copied into RNA.

Fig: Replication of DNA
The strand of DNA with polarity 3' → 5' that directs synthesis of RNA, is called template strand, noncoding strand or antisense strand. Another strand of DNA with polarity 5' → 3' is complementary to the template strand is called a coding strand, non template strand or sense strand. The template strand is transcribed into RNA. The primary mRNA produced contains codons with the same sequence as that of the coding strand exception of T for U and hence it is called the same.

Fig: Template and non template strand
Let’s take an example, to understand the process of transcription. Imagine a DNA with the sequence of 3’ - GAACTGCGA - 5’ (Refer the figure given below). Then the sequence of RNA will be 5’ - CUUGACGCU - 3’.

Fig: Sequence of RNA after transcription
Now if you notice, this sequence is similar to the sequence of non template strands. The only thing that differs is T is replaced by U in RNA. That’s amazing right? Hence strand 5’ → 3’ is also known as coding strand while 3’ →5’ strand is known as non coding strand.

Fig: Coding and non-coding strand
Reason for not processing both the strands of DNA
Now you will get a question, why does RNA polymerase not transcribe both the strands? Let’s try to understand this by considering what happens if both the strands were transcribed. If both strands act as a template, they would code for two RNA molecules with different sequences. This will result in the formation of two different proteins that can have a detrimental effect on the cell.

Fig: Transcription of both DNA strands
Secondly, the two RNA molecules produced would be complementary to each other, hence would form a double stranded RNA or RNA dimer. This would prevent RNA from being converted into protein, as the ribosomes cannot process double stranded mRNA.

Fig: RNA dimer
Transcription is selective
The entire molecule of DNA is not expressed in transcription. RNAs are synthesised only for some selected regions of DNA. Certain regions of DNA may not have any transcription at all. This happens due to the presence of some inbuilt signals in the cell.
Transcription unit
Transcription unit is defined as the segment of DNA between the sites of initiation and termination of transcription by RNA polymerase. There will be more than one gene that may reside in a transcription unit. The transcription unit mainly consists of three regions of DNA. They are structural genes, promoter region and terminator region.

Fig: Transcription unit
Gene
Gene has the nucleotide sequence which codes for RNA molecules. The final RNA molecule will have a sequence similar to the region. So a gene can be defined as the functional unit of inheritance.

Fig: Gene
Cistron
Genes are located on DNA and it is difficult to literally define a gene in terms of DNA sequence. DNA sequence coding for tRNA or rRNA molecules also defines a gene. Cistron is defined as a functional unit of gene, it is a segment of DNA coding for a polypeptide.

Fig: Structure of gene
Structural gene
The area of the template strand that is involved in transcription or formation of RNA is called the structural gene. There are two types of structural genes depending upon the number of cistrons present in it. They are monocistronic and polycistronic.
Fig: Types of structural genes
Monocistronic structural genes
These are made up of only one cistron. It carries information for synthesis of one polypeptide chain. It is mostly found in eukaryotes.
Polycistronic structural genes
These are made up of many cistrons. It carries information for synthesis of more than one polypeptide chain. They are mostly found in prokaryotes
Promoter
Promoter is a region of DNA molecules to which an RNA polymerase binds and initiates transcription. The promoter is located upstream (to the left) of the structural gene, i.e., towards the 5' end of the coding strand or 3' end of the template strand.
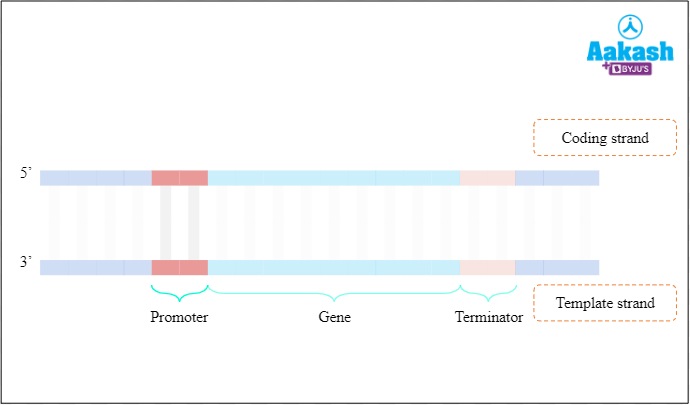
Fig: Promoter region
Location of promoter gene and terminator gene is always mentioned with reference to the coding strand.

Fig: Promoter and terminator
Promoter sites in prokaryotes
There are two promoter sites in prokaryotes which are commonly recognised by the sigma factor of the DNA dependent RNA polymerase.
TATA box
TATA box is a widely known promoter site. It is named as the TATA box is found to have the sequence TATAAT (six nucleotides). The area has a groove to which specific protein components can combine. It is located about 10 bases away (upstream) from the starting point of transcription. TATA box is also called Pribnow box (after its discoverer Pribnow) in prokaryotes.

Fig: TATAAT sequence
-35 sequence
This is the second recognition site in the promoter region of genes in prokaryotes. It contains a base sequence TTGACA which is located about 35 bases (upstream) away on the left side from the site of the starting point of transcription.
Promoter sites in eukaryotes
There are two promoter sites in eukaryotes which are commonly recognised by the DNA dependent RNA polymerase.
Hogness box
Hogness box (after its discoverer Hogness) in eukaryotes is the most common promoter site. Hogness box is located on the left about 25 nucleotides away from the starting point of initiation. Besides a promoter, eukaryotes also require an enhancer.

Fig: Names of TATA box
CAAT box
There exists another promoter site called CAAT box in eukaryotes. It is located 70 - 80 nucleotides upstream from the starting point of transcription. It possesses the sequence GGCCAATCT.
Enhancer
These can increase the gene expression by 100 fold. These enhancers bind to the transcription factors to form activators.
Terminator
Terminator region is present downstream or 3’ end of coding strand of gene. When RNA polymerase reaches this site, transcription stops and primary RNA transcript is released.
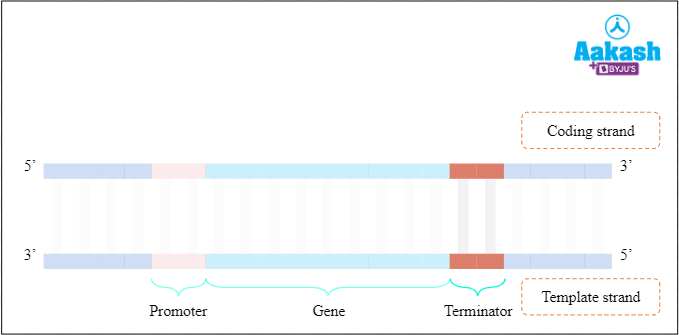
Fig: Terminator
Requirements for transcription
- Enzyme: DNA dependent RNA polymerase
- DNA template
- Four types of ribonucleotides triphosphates (ATP, CTP, GTP and UTP)
- Divalent metal ions Mg2+ or Mn2+ as a cofactor
DNA dependent RNA polymerase
The DNA dependent RNA polymerase differs in eukaryotes and prokaryotes. There is a single DNA dependent RNA polymerase in prokaryotes which synthesises all types of RNAs. Eukaryotes have three types of RNA polymerases and these are RNA polymerase I, RNA polymerase II and RNA polymerase III.
Functions of DNA dependent RNA polymerases in eukaryotes
Now let’s see what are the functions of these polymerases.
- RNA Polymerase I - It synthesises large ribosomal RNAs or rRNAs (28S, 18S and 5.8S) in the nucleolus.
- RNA Polymerase II - It synthesises mRNA, SnRNA (Small nuclear RNA) and hnRNA (heterogenous nuclear RNA).
- RNA Polymerase III - It synthesises tRNA and 5S rRNA (small ribosomal RNA) in the nucleolus.
Structure of DNA dependent RNA polymerase
It is a large complex consisting of 4 subunits which make the core enzyme and one σ (sigma) subunit and ρ subunit.

Fig: Parts of RNA polymerase
Core enzyme
It has four subunits as follows:
- Two ∝ Subunits : These bind to regulatory proteins.
- One β Subunit : It binds to the DNA template.
- One β’ Subuni : It binds to the ribonucleotides.
σ subunit and ρ subunit
Sigma subunit is for promoter recognition and initiation of transcription. Transcription termination is done by the Rho subunit.
Transcription in prokaryotes
What do you know about the DNA of bacteria? Yes, prokaryotes have circular DNA. So how does the transcription happen in the prokaryotes? Let’s check it out.
RNA synthesis in prokaryotes
In prokaryotes the structural genes are polycistronic and continuous. The process of transcription is completed in three steps. They are initiation, elongation, and termination.
Initiation
The first phase of transcription is initiation. As we mentioned earlier, transcription requires DNA as template and DNA dependent RNA polymerase. But to initiate the transcription, DNA dependent RNA polymerase requires one more element. It is the initiation factor or sigma factor. This factor directs the DNA dependent RNA polymerase towards the promoter region and helps in binding to the promoter to start transcription.
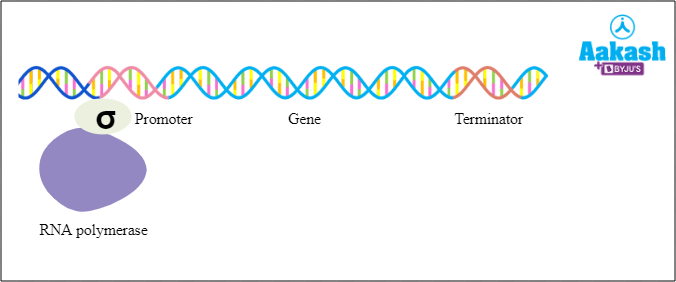

Fig: Transcription; Initiation
RNA polymerase and sigma factor moves forward towards the gene. The DNA dependent RNA polymerase now unwinds the DNA strands around the transcription start site.

Fig: RNA polymerase and sigma factor moving forward towards gene
RNA polymerase picks up the ribonucleotides and starts adding them with DNA as a template based on the complementarity rule. Here, in the image the lower strand acts as template strand and upper strand acts as coding strand.

Fig: RNA polymerase picks up the nucleotides and adding them with DNA as template
According to the rule of complementarity of bases, Cytosine always pairs with Guanine. In RNA Adenine base pairs with Uracil.

Fig: Base pairing
Let’s look at the sequence of coding strands. You can see that the coding strand has Adenine followed by Cytosine. Usually Thymine base pairs with Adenine. But in RNA, Uracil is present instead of Thymine, so RNA polymerase adds UTP over ATP. Similarly, RNA polymerase keeps adding ribonucleotides as per the base complementarity rule.
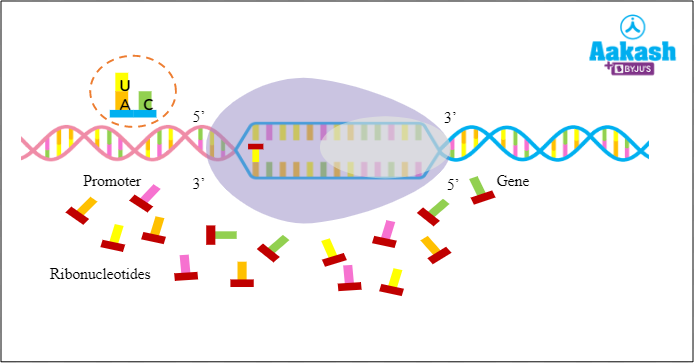
Fig: Polymerase adding UTP over ATP
Size of RNA eventually grows as and when new ribonucleotides are added. As the polymerase moves ahead, the free end of RNA is detached from the DNA template and the unwound DNA gets rewinded back.

Fig: Growing RNA detaching from the DNA template
After adding a few ribonucleotides, sigma factor is released from RNA polymerase. Release of sigma factor begins the next stage which is elongation.
Elongation
During elongation, RNA synthesis continues. For the addition of each ribonucleotide to the growing chain a pyrophosphate moiety is released. As DNA dependent RNA polymerase moves across, only a small part of RNA is attached to the template strand, the other end of growing RNA is free. Elongation continues without any interruption until the polymerase encounters the terminator region.

Fig: Elongation
When the elongation ends, the next phase of transcription begins which is termination.
Termination
Two types of termination are present in prokaryotes. These are the rho dependent termination and the rho independent termination.
Rho dependent termination
Termination begins when another factor known as termination factor or rho factor binds to RNA. It finally reaches the region where RNA is bound to the DNA template strand. After reaching this region, the rho factor unwinds the DNA-RNA complex. Thus causing the release of RNA from the DNA dependent RNA polymerase.

Fig: Rho factor initiating termination
Once the RNA is released, DNA dependent RNA polymerase and rho factor comes off of the DNA strand. DNA gets rewinded back into helical structure. This marks the end of RNA transcription.

Fig: RNA and Rho factor detaching from the DNA
Rho independent termination
In this model the termination is bought by the formation of hairpins in the newly synthesised RNA. This occurs due to the presence of palindromes. The presence of palindromes in the base sequence of DNA template creates palindromes in the primary RNA transcript. As a result of this the newly synthesised RNA folds to form hairpins due to complementary base pairing which causes termination of transcription.

Fig: Rho independent termination
Characteristic features of RNA synthesis in prokaryotes
The following are the major features of RNA synthesis in prokaryotes:
- mRNA does not require any processing to become active unlike in eukaryotes.
- Transcription and translation take place in the same compartment called cytosol as there is no separation of cytosol and nucleus.
- Many times the translation can begin much before the mRNA is fully transcribed. Hence the transcription and translation can occur simultaneously.
Transcription in eukaryotes
The transcription process in eukaryotes has the same steps as in prokaryotes but is a little more complex or elaborate. It occurs in G1 and G2 phases of cell cycle. There are only a few differences in them. Eukaryotic transcription occurs in the nucleus, chloroplast and mitochondria. But for prokaryotes it occurs in the cytoplasm as their DNA is in the cytoplasm. The steps of eukaryotic transcription also include initiation, elongation and termination. Now let's discuss some of the features of eukaryotic transcription.
Features of transcription eukaryotes
The following are the major features of transcription in eukaryotes:
- The eukaryotic transcriptional unit has only one gene (monocistronic).
- Initiation requires proteins called transcription factors.
- Sigma factor is absent in DNA dependent RNA polymerase.
- Do not require a primer to initiate RNA synthesis.
- RNAs are released and processed in the nucleus.
- Coupled transcription-translation is not possible as translation occurs in the cytosol.
- A greater part of products pass from the nucleus into the cytoplasm.
Like we discussed about DNA dependent RNA polymerase in prokaryotes, eukaryotes also have DNA dependent RNA polymerases with the same features. The only difference is that instead of one it has three RNA polymerases. But why do they have three RNA polymerases? In eukaryotes there are some additional RNAs. So depending on the types of RNA transcribed, they need more DNA dependent RNA polymerases.
So now we will get an idea about different types of RNAs.
Types of RNA
There are three major types of RNAs. These are mRNA, tRNA and rRNA. Let’s discuss more about it.
mRNA or messenger RNA
The RNAs which carry messages from DNA are mRNA. They are a template for protein synthesis. mRNAs can have one or more cistrons. Cistron means the nucleotide sequence which codes for a single protein.
Types of mRNAs
Based on the number of cistrons, mRNA can be classified as monocistronic or polycistronic.

Fig: Types of mRNA
Monocistronic mRNA
It contains a single cistron. So this would mean that monocistronic mRNA has information for synthesis of single protein only. It is found in eukaryotes.
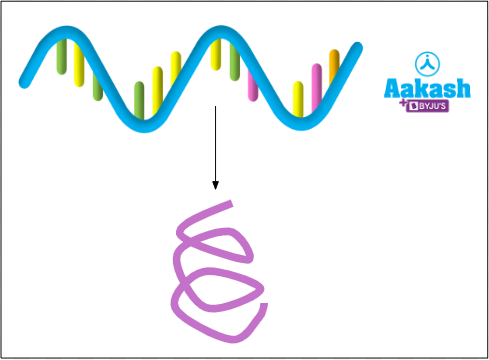
Fig: Monocistronic mRNA in eukaryotes
Polycistronic mRNA
It contains multiple cistrons. Polycistronic mRNA will have information which codes for more than one protein. It is found in prokaryotes.
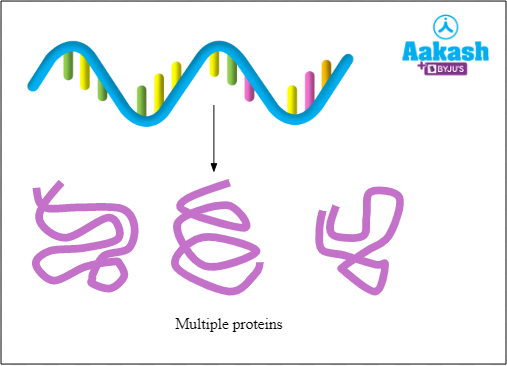
Fig: Polycistronic mRNA in prokaryotes
tRNA or transfer RNA
After the synthesis of tRNA, it folds itself to form clover shape. tRNA carries correct amino acids to the site of protein synthesis in the ribosome. All the tRNAs undergo post-transcriptional modifications. This includes trimming, converting the existing bases into the unusual ones, addition of CCA nucleotides to the 3’ terminal end of the tRNA.

Fig: tRNA
rRNA or ribosomal RNA
We know that ribosomes synthesise proteins and they are made up of RNA and proteins. The RNAs present in ribosomes are rRNAs. They help in catalysing protein synthesis. rRNA also undergoes post transcriptional modifications.
So now it is clear that there are different types of RNAs. So for transcribing different RNAs we need specific polymerases. Let’s find out which are the polymerases used for different RNAs.
Types of DNA dependent RNA polymerases
RNA polymerase I is used for transcribing rRNA (28S, 18S and 5.8S). RNA polymerase II is used for transcribing mRNA, SnRNA and hnRNA. RNA polymerase III is used for transcribing tRNA and 5S rRNA.
In addition to the above DNA dependent RNA polymerases, eukaryotes possess special DNA dependent RNA polymerases in the mitochondria. This resembles the prokaryotic DNA dependent RNA polymerases.
Process of eukaryotic transcription
The transcription process in eukaryotes is complex. It involves initiation, elongation and termination.
Initiation
The initiation step is divided into three stages as follows:
- Chromatin containing the promoter sequence is made accessible to the transcription machinery.
- There are some factors or proteins that are necessary for the DNA dependent RNA polymerase to bind the DNA. These proteins are called transcriptional factors or TF and they help in recognizing the promoter sequence in the transcription unit. These transcription factors include TFIID, TFIIA, TFIIB, TFIIF, TFIIE and TFIIH. These TFs bind to each other and to the enzyme DNA dependent RNA polymerase. Once the TF recognises the promoter region, the DNA dependent RNA polymerase binds to it and starts the transcription process.

Fig: Transcription factor on the DNA template
- Stimulation of the transcription by enhancers. This is followed by elongation and termination steps which finally results in the production of the primary RNA transcript.

Fig: RNA polymerase binds on the TF
Elongation
Elongation in eukaryotes is the same as that of prokaryotes except that it does not have a sigma factor in its DNA dependent RNA polymerase.
Suppose ATP is the first nucleotide, you know in RNA, Uracil is present instead of Thymine, so DNA dependent RNA polymerase adds UTP over ATP. If the next nucleotide on the coding strand is CTP, DNA dependent RNA polymerase picks up GTP and adds it next to UTP. If the third nucleotide in the coding strand is ATP, then the third nucleotide added on RNA will be UTP again. Similarly, DNA dependent RNA polymerase keeps adding nucleotides as per base complementarity. Size of RNA eventually grows as and when new nucleotides are added.

Fig: Elongation in eukaryotic transcription
Termination
In most cases in eukaryotes, there is a termination protein that stops the transcription process.
As you know after the termination, the RNA polymerase dissociates along with it the termination protein or TP also dissociates and primary RNA transcript is formed.

Fig: Termination by TP in eukaryotes
Post – transcriptional modifications
The nascent or newly formed RNA synthesised by RNA polymerase II is called hnRNA (heterogenous nuclear RNA) or primary RNA transcript. It is non-functional and without modifications, it cannot participate in protein synthesis. It needs to undergo modifications to form functional mRNA, rRNA or tRNA. So it will undergo further processing to mature in the nucleus. The primary transcript is converted into functional mRNA after post-transcriptional processing which involves the following steps:
- Splicing
- Capping
- Tailing
- Ligation

Fig: Steps of Post transcriptional modification
Splicing
Splicing is the process of removal of extra segments from the RNA transcript. But what exactly is extra in the transcript? The primary transcript contains unwanted base sequences called introns and useful base sequences called exons. Introns are non coding, non functional or intervening sequences in the RNA. Exons are the coding or functional sequence in the RNA that codes for proteins.

Fig: Primary transcript
Removal of introns through cutting and joining of exons in a defined order is called splicing. Introns are removed with the help of snRNPs (small nuclear ribonucleoprotein particles). It is assembled from proteins and smaller nuclear RNA (snRNA) molecules that recognise splice sites in the pre-mRNA sequence. The splicing reaction is catalysed by a large protein complex called the spliceosome. Spliceosomes represent the snRNP association with the hnRNA at the exon-intron junction. Removal of introns and joining of exons leads to the bending of the introns and bringing exons close to each other.

Fig: Splicing using spliceosome
The ends of the adjacent exons are joined together by ligase enzymes. The snRNPs are formed by association of proteins with small nuclear RNAs. It probably represents an ancient feature of the genome.

Fig: Joining by ligase enzyme
The presence of introns is reminiscent of antiquity and the process of splicing represents the dominance of the RNA-world. After splicing the next step is capping.
Capping
Capping is the addition of nucleotide like methylated guanosine triphosphate (mGppp) by the unusual 5’ → 5’ linkage on the 5’ end. This structure is required for translation and stabilisation of the mRNA structure.

Fig: Capping
Tailing
In tailing, adenylate residues (200-300) called poly-A tail are added at 3'-end in a template independent manner. This is required for the stabilisation of the mRNA structure. Poly-A tails get reduced as the mRNA enters the cytosol.
Fig: Tailing
After all these modifications finally the hnRNA is now a functional mRNA. Following this the mRNA leaves the eukaryotic nucleus. The transcript is transported out of the nucleus to the cytosol for translation.

Fig: Mature mRNA
An advantage of post transcriptional modification is that the cap protects the 5' end of the primary RNA transcript from attack by enzymes like ribonucleases. Modifications also help the RNA molecule to be recognised by molecules that mediate RNA translation for the production of proteins.

Fig: Modification protecting the mRNA from enzyme attack
Reverse transcription
In some viruses like retroviruses, the RNA is transcribed into DNA. This process of transcribing RNA into DNA is called reverse transcription. The enzyme responsible for the synthesis of DNA from RNA template is called reverse transcriptase. HIV is an example for the virus which has reverse transcriptase enzyme.

Fig: Reverse transcription
Central dogma reverse
Howard Temin and David Baltimore discovered reverse transcriptase enzymes in 1970. This enabled information to flow in the direction RNA → DNA. This is called central dogma reverse.

Fig: Central dogma reverse
Application of central dogma reverse
This principle is applied in the production of cDNA. For example when DNA with the gene of interest needs to be inserted into prokaryotes, as they have no machinery to process the primary RNA transcript, mostly cDNAs are used.
Synthesis of cDNA from mRNA
The DNA expresses the genetic information in the form of RNA. The mRNA determines the amino acid sequence in a protein. The mRNA can also be utilised as a template for the synthesis of double stranded complementary DNA (cDNA) by using the enzyme reverse transcriptase. This cDNA can be used as a probe to identify the sequence of DNA in genes.
Significances of transcription and post transcriptional modifications
The following are the major significances of transcription and post transcriptional
modifications:
- The transfer of information from DNA to RNA is only possible through transcription.
- The non-coding segments of RNA transcript (intron) are removed during splicing.
- Capping protects the 5' end of the primary RNA transcript from attack by enzymes like ribonucleases.
- Tailing provides stabilisation to the mature RNA.
- A well modified RNA is easily identified by the molecules which helps in the translation to make proteins.
- Post transcriptional modifications allow the transfer of mature RNAs to the cytosol.
- Faulty splicing can result in nonfunctional mRNAs. For example, in beta thalassemia, lack of synthesis of the beta chain of haemoglobin results due to a nucleotide change in the exon-intron junction.
Practice Problems
Q1. A geneticist finds the sequences of the coding strands in transcription units as follows:
(a) 5'-TGAACTGTAGCATGC-3'
(b) 5'-TGATACTGGTACATTC-3'
Find out the correct sequences of the mRNAs transcribed from them.
A. (a) 5'-UGAACUGUAGCAUGC-3'
(b) 5'-UGAUACUGGUACAUUC-3'
B. (a) 3'-CGUACGAUGACAAGU-5'
(b) 3'-TGATACTGGTACATTC-5'
C. (a) 3'-CGUACGAUGUCAAGU-5'
(b) 3’-TGATACUGGTACAUTC-5'
D. (a) 5'-UGACUGUAGCUUGCC-3'
(b) 5'-TGATACTGGTACATTC-3'
Solution: If the coding strand in a transcription unit is having the following sequence:
5'-TGAACTGTAGCATGC-3', then, the sequence of mRNA transcribed is the same as the coding strand of DNA. However, in the newly formed RNA, Thymine is replaced by Uracil. Hence, the sequence of mRNA will be 5'-UGAACUGUAGCAUGC-3'.
Similarly, if the coding strand in a transcription unit is having the following sequence:
5'-TGATACTGGTACATTC-3', then the sequence of mRNA will be 5'-UGAUACUGGUACAUUC-3'. Hence the correct option is a.
Q2. The given diagram represents the transcription unit. Select the correct labelings regarding it.

- a. A - Terminator, B - Promoter, C - Template strand, D - Coding Strand
b. A - Promoter, B - Terminator, C - Template strand, D - Coding strand
c. A - Promoter, B - Terminator, C - Coding strand, D - Template strand
d. A - Promoter, B - Template strand, C - Terminator, D - Coding strand
Solution: ‘A’ is the promoter which is located upstream (5’ end) of the coding strand. The process of transcription initiates here with the help of the DNA dependent RNA polymerase enzyme. 'B' is the terminator, the process of transcription terminates here. It is located towards the 3' end (downstream) of the coding strand. All the reference points while explaining the transcription unit are made with respect to the coding strand. 'C ' is the template strand. It is a strand of DNA which has the polarity of 3’ → 5’. It acts as a template strand because transcription takes place in the 5' → 3’ direction. 'D' is the coding strand. It is a 5’ → 3’ strand of DNA. It is called coding strand because its nitrogenous base sequences are the same as the transcribed RNA, except for Uracil. In place of the Uracil of RNA, the coding strand of DNA has Thymine. Hence the correct option is b.
Q3. Assertion: Template or antisense strand, having 3 '→ 5' polarity takes part in transcription.
Reason: Nontemplate or sense strand, having 5' → 3' polarity, does not take part in transcription.
A. Both the assertion and the reason are true and the reason is the correct explanation of the assertion.
B. Both the assertion and the reason are true but the reason is not the correct explanation of the assertion.
C. The assertion is true but the reason is false.
D. Both the assertion and the reason are false.
Solution: The process of copying genetic information from one of the two strands of DNA (template strand) into RNA is called transcription. RNA polymerase transcribes RNA in 5' → 3' direction only. That is why the DNA strand having polarity 3' → 5' acts as a template strand or antisense strand and takes part in transcription. The other DNA strand having polarity 5' → 3' acts as a non-template strand or sense strand and does not take part in transcription. Therefore, both the assertion and the reason are true but the reason is not the correct explanation of the assertion. Hence the correct option is b.
Q4. Match column I with column II and choose the correct option.
|
Column I |
Column II |
|
|
|
2. Rho factor |
|
|
3. Alpha unit |
|
A. 1 - A , 2 - B , 3 - C
B. 1 - C, 2 - A, 3 - B
C. 1 - B, 2 - C, 3 - A
D. 1 - C, 2 - B, 3 - A
Solution: The RNA polymerase enzyme’s ‘core’ consists of 2-alpha subunits, 1-beta subunit, 1 β’ subunit, 1 omega subunit. RNA polymerase ‘core’ and the sigma factor interact to form RNA polymerase ‘holoenzyme’. A sigma factor is a protein needed only for initiation of RNA synthesis in bacteria. It is also called the specificity factor. Sigma factors provide promoter recognition specificity to the DNA dependent RNA polymerase and contribute to DNA strand separation. They then dissociate from the RNA polymerase core enzyme following transcription initiation. Rho (ρ) is an ATP-dependent RNA-stimulated helicase that disrupts the initial RNA-DNA complex and hence serves as a transcription termination factor. Hence the correct option is b.
Q5. Select the correct statements regarding transcription in prokaryotes and eukaryotes.
1. Capping and tailing takes place in prokaryotes.
2. Splicing takes place only in eukaryotes.
3. Transcription takes place in the cytoplasm in prokaryotes.
4. Prokaryotes have introns.
a. 1 and 2
b. 2 and 4
c. 2 and 3
d. 3 and 4
Solution: Transcription takes place in the nucleus in a eukaryotic cell. The prokaryotic cell lacks a well-defined nucleus and the genetic material is found in the cytoplasm. Thus, transcription in prokaryotes takes place in the cytoplasm. Transcription is different in eukaryotes as compared to prokaryotes as they have more complex cellular organisation. The primary transcript (hnRNA) formed after transcription in eukaryotes contains coding (exons) as well as non-coding (introns) sequences. The primary transcript of eukaryotes undergoes the following post-transcriptional changes:
- Capping - The addition of methyl guanosine triphosphate to the 5’ end.
- Tailing - The addition of adenylate residues at the 3’ end.
- Splicing - The removal of introns and linking of the exons. Hence the correct option is c.
FAQs
Q1. Why is mRNA not processed in prokaryotes?
Answer: In prokaryotes the transcription is taking place in the cytoplasm itself. So there's a chance for the mRNA to get degraded very rapidly from 5’ end by the cytoplasmic enzymes. Hence the translation will be started in the mRNA even before the transcription ends by the assembly of ribosomes on the first cistron of mRNA. So we can say that the transcription in prokaryotes is coupled with translation. As the mRNA does not need to cross the nuclear membrane barrier, capping and tailing is not required. Introns are absent in the mRNA of prokaryotes and hence the splicing is not required.
Q2. How is alternative splicing different from splicing?
Answer: Splicing is the process of splicing exons of primary transcript of mRNA. But the alternative splicing is the splicing of different combinations of exons of the same gene to get a particular protein from the primary transcript.
Q3. Why does tRNA have a clover leaf structure?
Answer: Clover leaf structure is the secondary structure of tRNA. The presence of regions of self-complementarity within tRNA creates the typical cloverleaf - shaped structure. This produces four stems (also called arms) and three loops.
Q4. What are mRNA vaccines?
Answer: mRNA vaccines are the piece of mRNA that is corresponding to the viral protein that causes infection. When this mRNA enters our body, the body cells will produce the viral protein. This results in the production of antibodies against this mRNA by considering it as a foreign particle. These antibodies remain in the body and prevent further infection from the particular virus.
YOUTUBE LINK: Need to be created
Related topics
|
The structure of DNA, Practice Problems and FAQs |
|
Gene regulation in prokaryotes, lac Operon, Practice Problems, FAQs |
|
DNA fingerprinting: Steps and Applications, Practice Problems and FAQs |


